Multiplex Tissue Imaging
Protein-Level Biomarker Discovery
Tumor biology is more than DNA and RNA. To understand spatial interactions between tumor and immune cells, LIDE offers next-generation tissue imaging platforms:
- 40+ validated antibody panels (OncoVee® Toolbox)
- Simultaneous detection of dozens of protein markers per FFPE slide
- Enables visualization of immune infiltration, signaling pathways, and microenvironment heterogeneity
- Example: Distinguishing responders vs. non-responders in checkpoint inhibitor studies
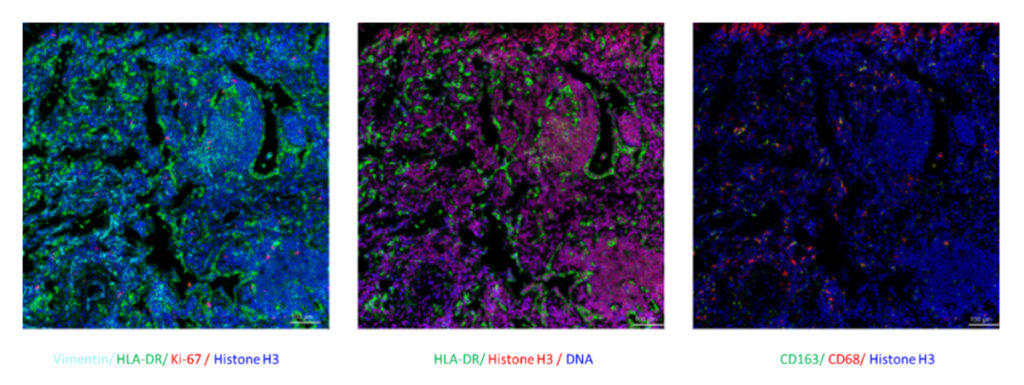
In addition, IMC is performed on FFPE tissue sections, so it can be applied to the retrospective research of disease and drug development using a limited number of pathological sections.
- Simultaneous detection of multiple proteins on a single tissue section
- Compatible with clinical workflow and FFPE archives
- Example markers: CD20, CD3, CD68, Ki67, E-cadherin
LIDE has developed a validated panel of IMC antibodies for leading oncology research targets, ready to use in analysis or customized according to need.
| Category | Examples |
|---|---|
| Cell Identification | Lineage and cell-specific markers to define tumor vs. stromal vs. immune cells |
| Tissue Histological Landmarks | Structural and microenvironmental features |
| Cancer Markers | Tumor-specific markers and oncogenic drivers |
| Major Immune Populations | T cells, B cells, NK cells, macrophages, dendritic cells |
| Cytokines | IL-family, TNF, IFN pathways |
| Granzyme B (GzmB) | Effector function marker for cytotoxic activity |
| Immune Checkpoints | PD-1, PD-L1, CTLA-4, TIM-3, LAG-3, TIGIT |
| Cell Proliferation & Death | Ki-67, cleaved caspase-3, apoptosis and proliferation markers |
- Companion diagnostic development
- Biomarker-driven patient stratification
- Mapping tumor–immune cell interactions
One of most valuable IMC advancements is that single-cell features can be computationally segmented and clusters of co-expression demarcated to enable downstream investigation of cell subpopulations and