CR (Conditionally Reprogrammed) Cells
Conditionally Reprogrammed (CR) Cells offer a transformative approach to in vitro modeling-forging a bridge between conventional cell lines and patient-derived in vivo models. LIDE’s CR platform enables rapid expansion of patient tumor tissues while preserving their original genetic, histological, and functional signatures.
Why CR Cells Matter:
| Benefit | Details |
|---|---|
| Preserves Tumor Integrity | Maintains both inter- and intra-tumoral heterogeneity-unlike immortalized cell lines |
| High Take Rate | Cultures can be established from nearly 90% of tissue samples without genetic manipulation |
| Genetic Fidelity | CR cell models retain key mutations (e.g. KRAS, TP53, SMAD4), matching original tumor profiles |
LIDE’s CR Cell Capabilities: 100+ CR lines derived from patient tumors, many matching LIDE’s PDX models and enriched for drug resistance or rare genotypes
| Sample Source | Tumor Type | #CRCs |
|---|---|---|
| PDX | Lung Cancer | 12 |
| Breast Cancer | 2 | |
| Melanoma | 1 | |
| Pancreatic Cancer | 18 | |
| Gastric Cancer | 2 | |
| Intestinal Cancer | 1 | |
| Esophageal Cancer | 4 | |
| Liver Cancer | 14 | |
| Cholangiocarcinoma | 10 | |
| Giloma | 4 | |
| Osteosarcoma | 1 | |
| Clinical Biopsy | Cardiac Cancer | 1 |
| Colorectal Cancer | 1 | |
| Lung Cancer | 2 | |
| Breast Cancer | 1 | |
| Clinical Puncture | Lung Cancer | 2 |
| Malignant Thoracic Peritoneal Mesothelioma | 1 | |
| Clinical Hydrothorax | Lung Cancer | 2 |
| Sample Source | Tumor Type | #CRCs |
|---|---|---|
| Clinical Surgery | Gallbladder Carcinoma | 4 |
| Cholangiocarcinoma | 1 | |
| Osteosarcoma | 1 | |
| Glioma | 3 | |
| Intestinal Cancer | 1 | |
| Oral Squamous Cell Carcinoma | 1 | |
| Lung Cancer | 3 | |
| Melanoma | 1 | |
| Oral Floor Carcinoma | 1 | |
| Renal Cancer | 2 | |
| Breast Cancer | 1 | |
| Esophageal Cancer | 1 | |
| Ovarian Cancer | 1 | |
| Gastric Cancer | 1 | |
| Pancreatic Cancer | 3 | |
| Colorectal Cancer | 1 | |
| Glioma | 3 |
| Tumor Type | Drug Resistance | Gene Alteration |
|---|---|---|
| NSCLC | Erlotinib Osimertinib Cetuximab EAI045 Cetuximab+EAI045 | EGFR: exon19del/L858R/T790M/C797S KIF5B-RET fusion EML4-ALK fusion ROS1: ROS1-RiLPL2/- MAP3K5-ROS1 fusion |
| BC | Primary CDK4/6 Inhibitor | |
| Melanoma | Imatinib Paclitaxel |
Validated Predictive Power
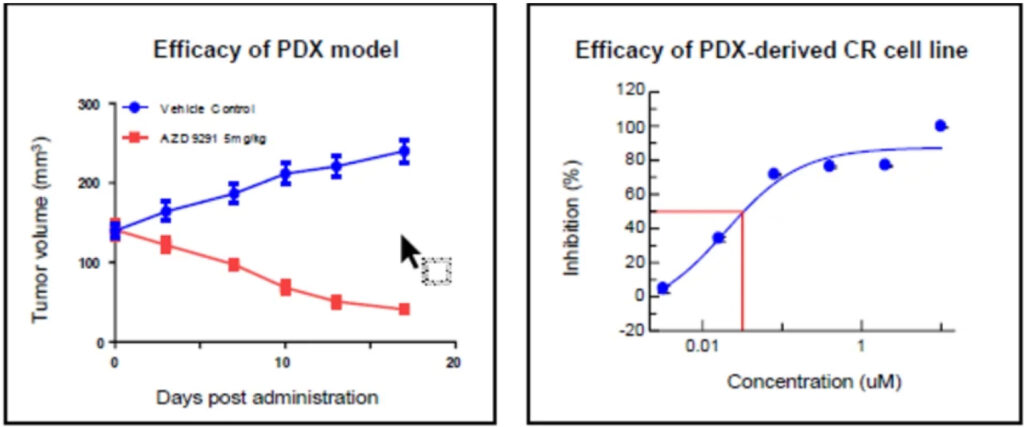
LIDE has confirmed that CR lines reflect PDX drug response profiles. For example, CR line with 19del/T790M/C797S mutation showed matching drug sensitivity (IC₅₀ ~0.03 µM or ~2.01 µM) to its parental PDX model. This alignment confirms both accuracy and translational value.
| Model ID | EGFR Status |
|---|---|
| LD1-0025-200636 | WT |
| LD1-0006-215676 | L858R/T790M |
| LD1-0025-200717 | 19del/T790M/C797S |
LD1-0006-215676
EGFR-L858R/T790M
LD1-0025-200717
EGFR 19del/T790M/C797S
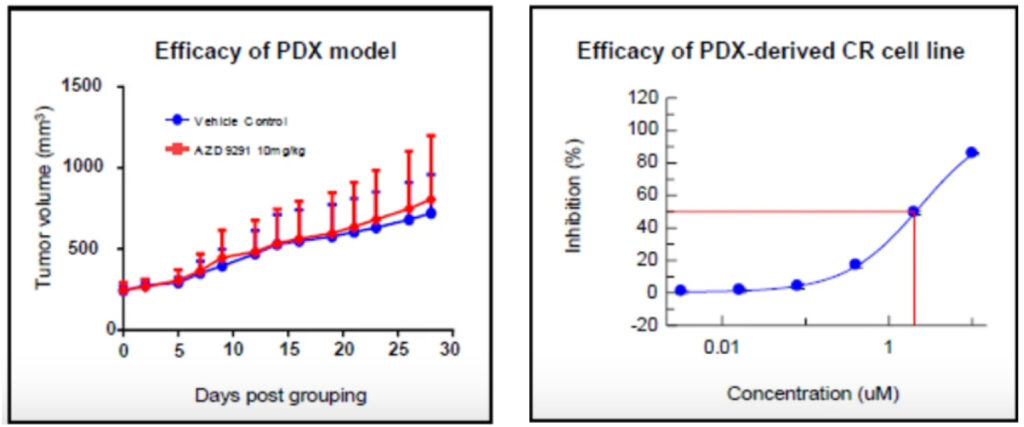
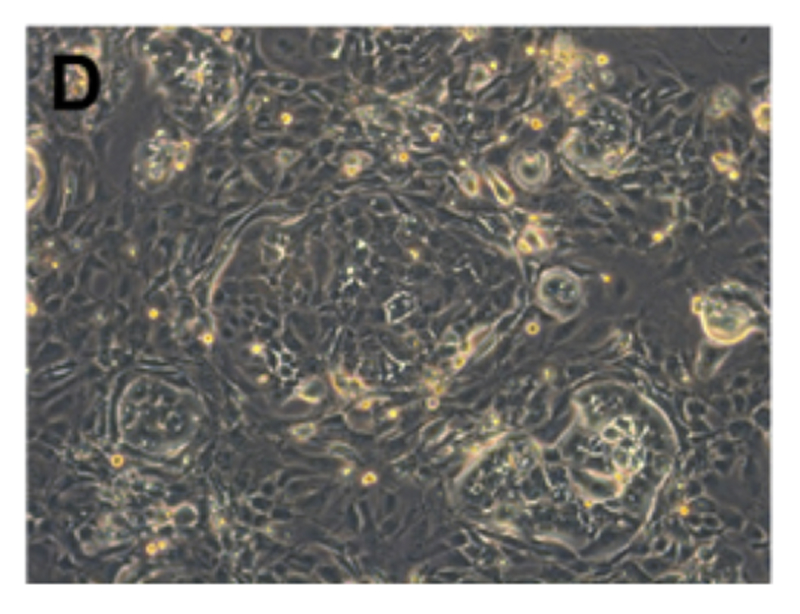
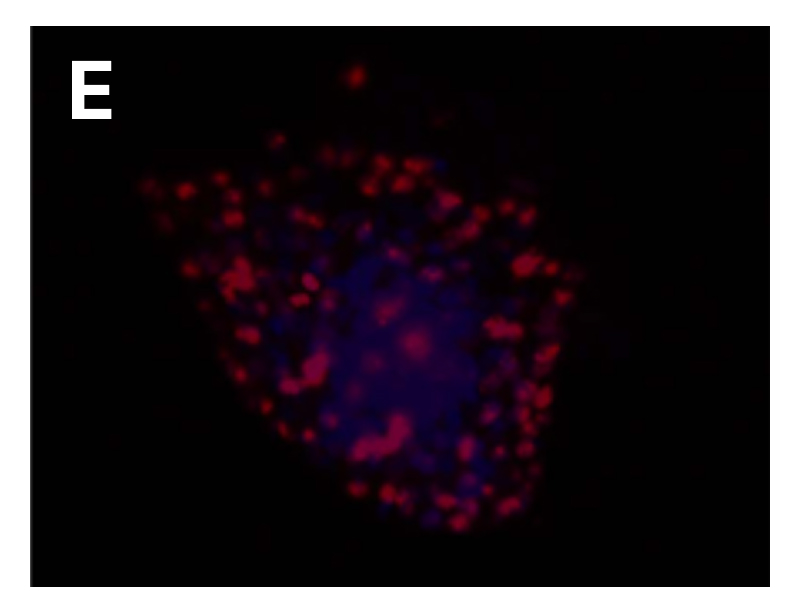
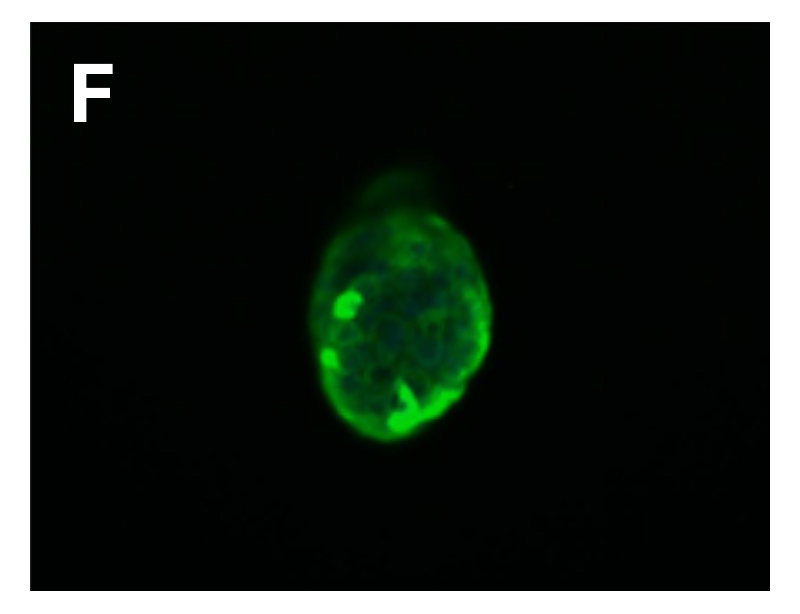
Fig. Morphology of EGFR 19del/T790M /C797S PDX derived cell line (D); Ki67 (E) and Pan-CK (F) staining of 19del/T790M /C797S PDX matching cell derived tumor sphere.
CR cultures support several applications:
- Early Drug Screening – Perform assays using patient-relevant biology.
- MiniPDX® Pairing – Scale in vivo insight from CR validation quickly.
- Targeted CDX Modeling – Seamlessly translate in vitro findings to in vivo systems.
- Biomarker Discovery – Link response signatures to CR cell phenotypes.
- Living Biobank Creation – Build sustained repositories from limited sample sources.
Additionally, LIDE has 220+ commercially available cancer cell lines already in house for similar 2D applications.